הפקולטה למדע והנדסה של חומרים
מטרות הפקולטה למדע והנדסה של חומרים בטכניון הינן להוביל את חזית המחקר בתחומים שונים במדע והנדסת חומרים, לשמש כמרכז לאומי ללימוד ומחקר בתחומי מדע והנדסת החומרים ולהכשיר מדענים ומהנדסים ברמה עולמית.
מטרות הפקולטה למדע והנדסה של חומרים בטכניון הינן להוביל את חזית המחקר בתחומים שונים במדע והנדסת חומרים, לשמש כמרכז לאומי ללימוד ומחקר בתחומי מדע והנדסת החומרים ולהכשיר מדענים ומהנדסים ברמה עולמית.

מהי הנדסת חומרים, מהן תוכניות הלימוד

חדשות אירועים וסמינרים

קבוצות מחקר, מחקרים ומעבדות

מערכת רישום לשימוש במיכשור פקולטי
הפקולטה למדע והנדסה של חומרים מציעה מספר תוכניות לימוד לתואר מוסמך במדעים (B.Sc)
תוכנית הלימודים לתואר ראשון בהנדסת חומרים, מעניקה לבוגריה תואר מוסמך למדעים (B.Sc) בהנדסת חומרים.
תוכנית לימודים משולבת לתואר ראשון בהנדסת חומרים ופיזיקה, מעניקה לבוגריה תואר מוסמך למדעים (B.Sc) בהנדסת חומרים ופיזיקה.
תוכנית הלימודים המשולבת לתואר ראשון בהנדסת חומרים וכימיה, מעניקה לבוגריה תואר מוסמך למדעים (B.Sc) בהנדסת חומרים וכימיה.
תוכנית הלימודים המשולבת לתואר ראשון בהנדסת חומרים וביולוגיה, מעניקה לבוגריה תואר מוסמך למדעים (B.Sc.) בהנדסת חומרים וביולוגיה.
מסלול למצטיינים במסגרת העתודה האקדמית של צה"ל בלימודי הנדסה, המקנה לבוגריו תואר ראשון (B.Sc) ושני (M.Sc) בהנדסת חומרים כולל תזה.
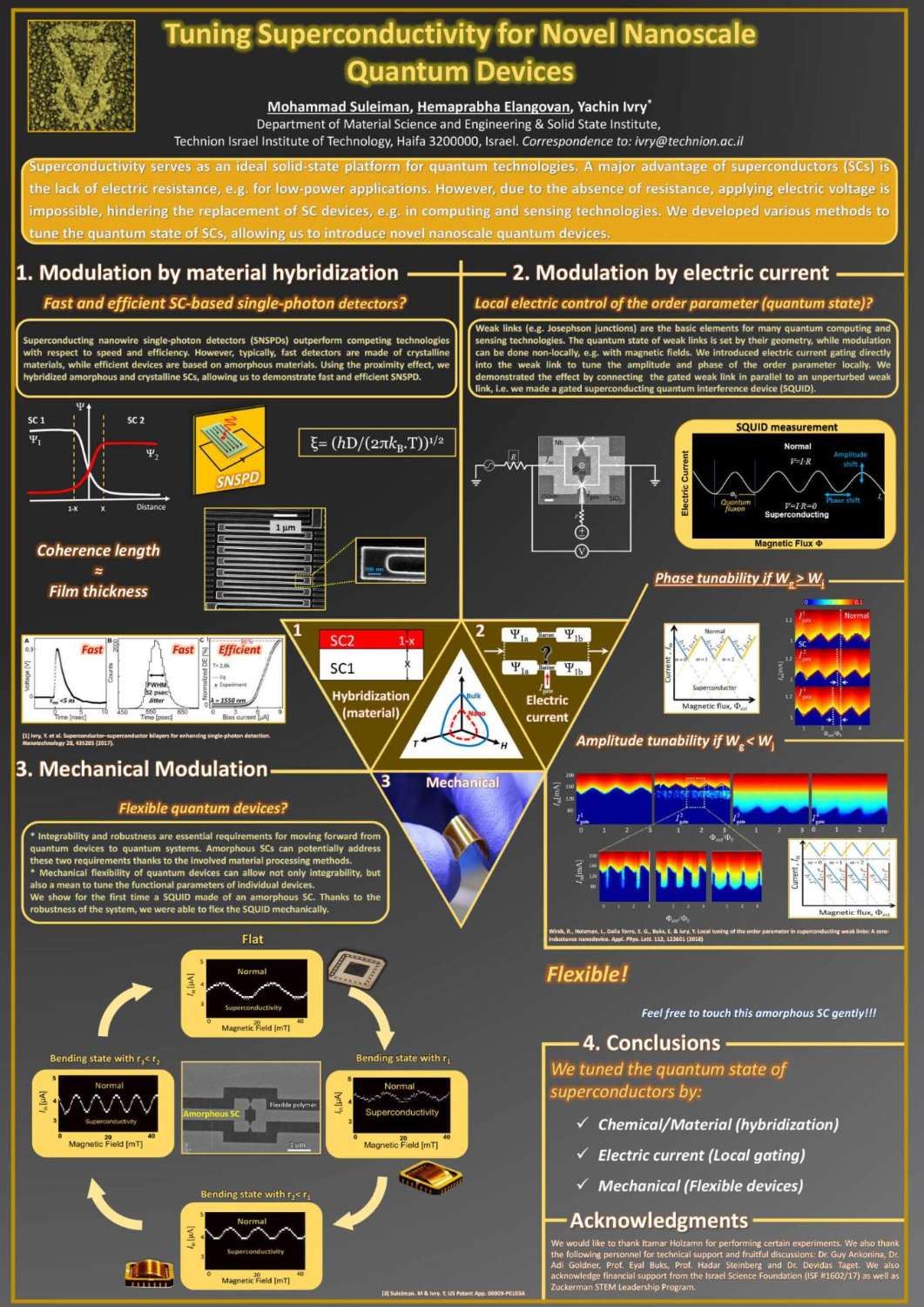
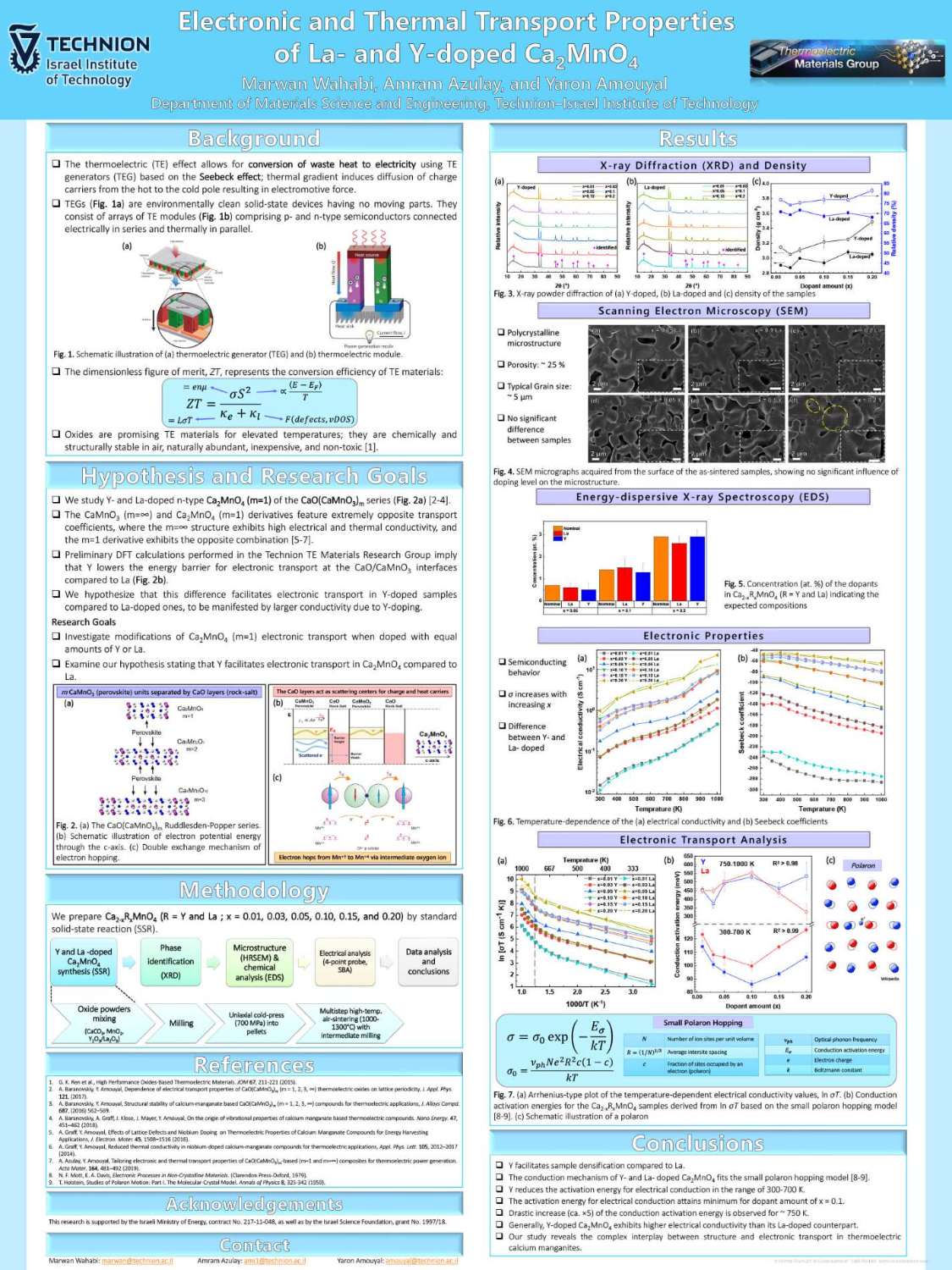
תצוגת פוסטרים של המשתלמים לתארים מתקדמים בפקולטה למדע והנדסה של חומרים


בוגרי הפקולטה למדע והנדסה של חומרים מספרים על חווית הלימוד בפקולטה ועל השתלבותם בתעשייה כמובילים וכמומחים בתחום הנדסת החומרים.